-
Notifications
You must be signed in to change notification settings - Fork 2
References
If you're confident your mapping reference (.fasta) will work with Castanet, start a Castanet run using this as input to the "RefStem" argument. You may leave the "MappingRefTable" argument as an empty string.
If you need to make a Castanet-compatible mapping reference, please read the following. It will guide you to generating a reference description file (.csv), which you can manually edit and input to Castanet runs as the "MappingRefTable" argument, alongside the "RefStem", to control how the probetype aggregation works.
Constructing a mapping reference (.fasta file) is the most important step in designing a Castanet run. This is because Castanet aggregates reads to probe sets (/organisms) through parsing (interpreting) the headers (">xyz") in your mapping reference file.
The two rules for how Castanet aggregates are as follows.
- Everything before the first underscore is treated as your "probetype", i.e. all reads mapped to references with the same text before an underscore get grouped together. The probetype field is mandatory.
- Within the "probetype", everything before the first hyphen is interpreted as the "organism". The organism field isn't essential, and if you have no hyphens in the probetype, the organism field will be set to the same value as the probetype.
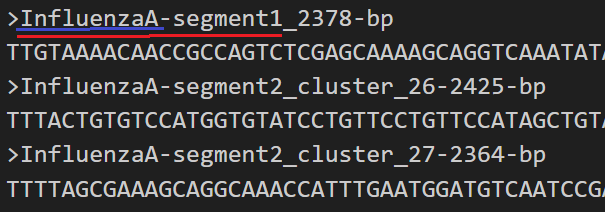
E.g in the above example, sequences mapped to top reference will aggregate to the probetype "InfluenzaA-segment1". Sequences mapped to the bottom two will all aggregate to "InfluenzaA-segment2". All reads mapped to any of these sequences will be labelled as organism "InfluenzaA".
Try to keep your headers to a sensible length, with no spaces, no delimiters other than underscores or hyphens, and no unprintable characters. This is a requirement because Castanet calls several command line tools that need to parse these headers.
Castanet has systems built in to either manually correct errors in headers, or otherwise inform users that their input are invalid, but as there are an almost infinite number of variations in headers, don't assume that the software is perfect at doing this!
If you're comfortable bulk-editing your mapping reference file using regular expressions, you're free to do so. Simply specify your mapping reference file as the "RefStem" argument, and leave the "MappingRefTable" argument as an empty string.
Castanet has a utility function to help design your mapping reference:
- API: /convert_mapping_reference/
- CLI:
$ python3 -m app.convert_mapping_reference --in_file XXX.fasta --out_file YYY.csv
This function loads a fasta file containing mapping references, and attempts to separate your headers into organism and probetype columns in a .csv file. You can then manually edit these fields, and input them to your run alongside your mapping reference file. Castanet will obey aggregation rules set in the .csv, rather than in the .fasta.
Example workflow:
- Source a mapping reference .fasta file. The example given below is an example of non-compatible mapping reference headers that can be converted using this function.
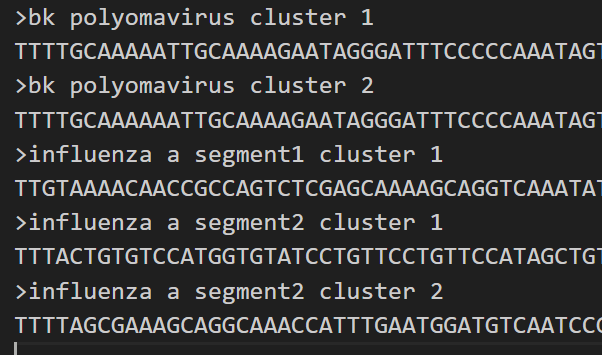
-
Run the mapping reference converter.
- In the API, find the '/convert_mapping_ref/' endpoint, specify your input fasta file as "InFile" and where to save your output csv as "OutFile".
- In the CLI, run
$ python3 -m app.convert_mapping_reference --in_file XXX.fasta --out_file YYY.csv
The example below shows the .csv output from the .fasta file shown in step 1. As there were no hyphens or underscores, the converter was unable to accurately determine the organism or virus name from the header. Headers where no probetype can be determined are automatically labelled "unaggregated", to make it easier to find them.
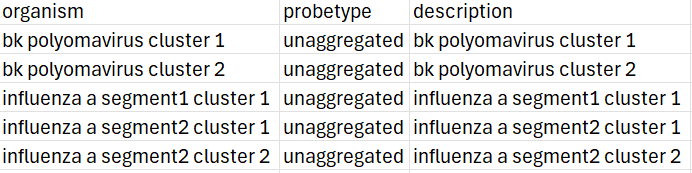

- Manually edit the file to show the correct organism and probetype (keeping in mind to avoid spaces, delimiters etc.)

- Start your Castanet run, specifying your original mapping reference file in the "RefStem" field, and your edited csv file as the "MappingRefTable" field. The output from a representative Castanet run is shown below (XXX_depth.csv), showing how reads were aggregated to each probetype.

Instructions are identical to those for API users, with the exception that the CLI wrapper can be accessed with:
$ python3 -m app.convert_mapping_ref -InFile ./my_mapping_ref.fasta -OutFile ./my_mapping_ref_table.csv