-
Notifications
You must be signed in to change notification settings - Fork 2
Description
Hi @RaoulWolf
Paul has added some new standardize options to PUG REST that is really going to help us deal with mixtures/components/neutralizing a bit better. As yet undocumented their side but here's an example:
- SMILES (an example not in PubChem):
CC(=O)OC1=C(Cl)C=CC=C1C(=O)[O-].CC(=O)OC1=C(Cl)C=CC=C1C(=O)[O-].[Ca+2] - Query (SMILES encoded): https://pubchem.ncbi.nlm.nih.gov/rest/pug/standardize/smiles/SDF?smiles=CC%28%3DO%29OC1%3DC%28Cl%29C%3DCC%3DC1C%28%3DO%29%5BO-%5D.CC%28%3DO%29OC1%3DC%28Cl%29C%3DCC%3DC1C%28%3DO%29%5BO-%5D.%5BCa%2B2%5D
... will get you an SDF with following notes (quoting a lot from Paul):
- CID will be zero if the structure isn’t in PubChem;
- the compound type is 1 for the full standardized record, and 2 for the components;
- the parent is indicated by InChIKey, and the parent – if any - will always be either self or one of the components;
- small subset of properties (SMILES, InChI, InChIKey etc) as it's computationally intensive (other properties should be retrieved/calculated via other routes);
- input as InChI, SDF is also possible.
So with this very basic code I did up a test set based on some examples from @hansarp and @sarahehaleZeroPM
UBAPMT_TestSet.csv
test_set <- read.csv("UBAPMT_TestSet.csv", stringsAsFactors = F)
test_set$PC_URL <- ""
i <- 1
for (i in 1:length(test_set$Order)) {
# for (i in c(1,5,11,13,26)) { # easy examples
# for (i in c(29:38,43)) { # trickier examples
SMILES <- test_set$SMILES_in[i]
PC_URL <- paste0("https://pubchem.ncbi.nlm.nih.gov/rest/pug/standardize/smiles/SDF?smiles=",
URLencode(SMILES, reserved = T))
test_set$PC_URL[i] <- PC_URL
download.file(PC_URL,paste0(test_set$InChIKey[i],".sdf"))
}
write.csv(test_set,"UBAPMT_TestSet_wURL.csv",row.names=F)
So, you can get lots of SDFs ... which contain what we need ...
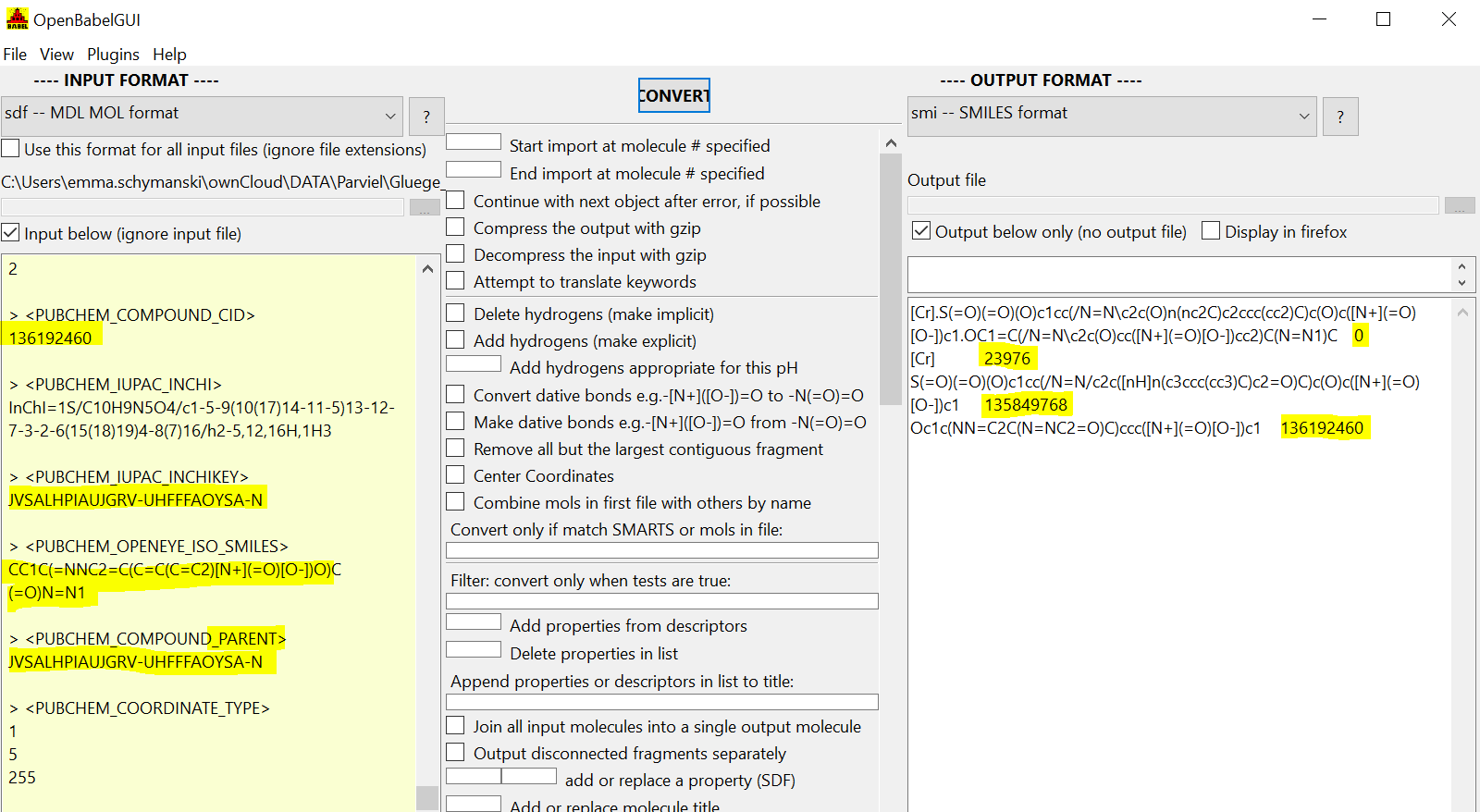
For example SEJWJMPJJMJBJG-VRQREAPISA-I.sdf (can't upload SDFs so this hyperlinks the query URL) you get:
- the entry you input at the top (not in PubChem, so CID=0)
- the components in the next three rows on the SMILES side with their CIDs (if available)
- the properties, including Parent InChIKey in the SDF tags (see left)
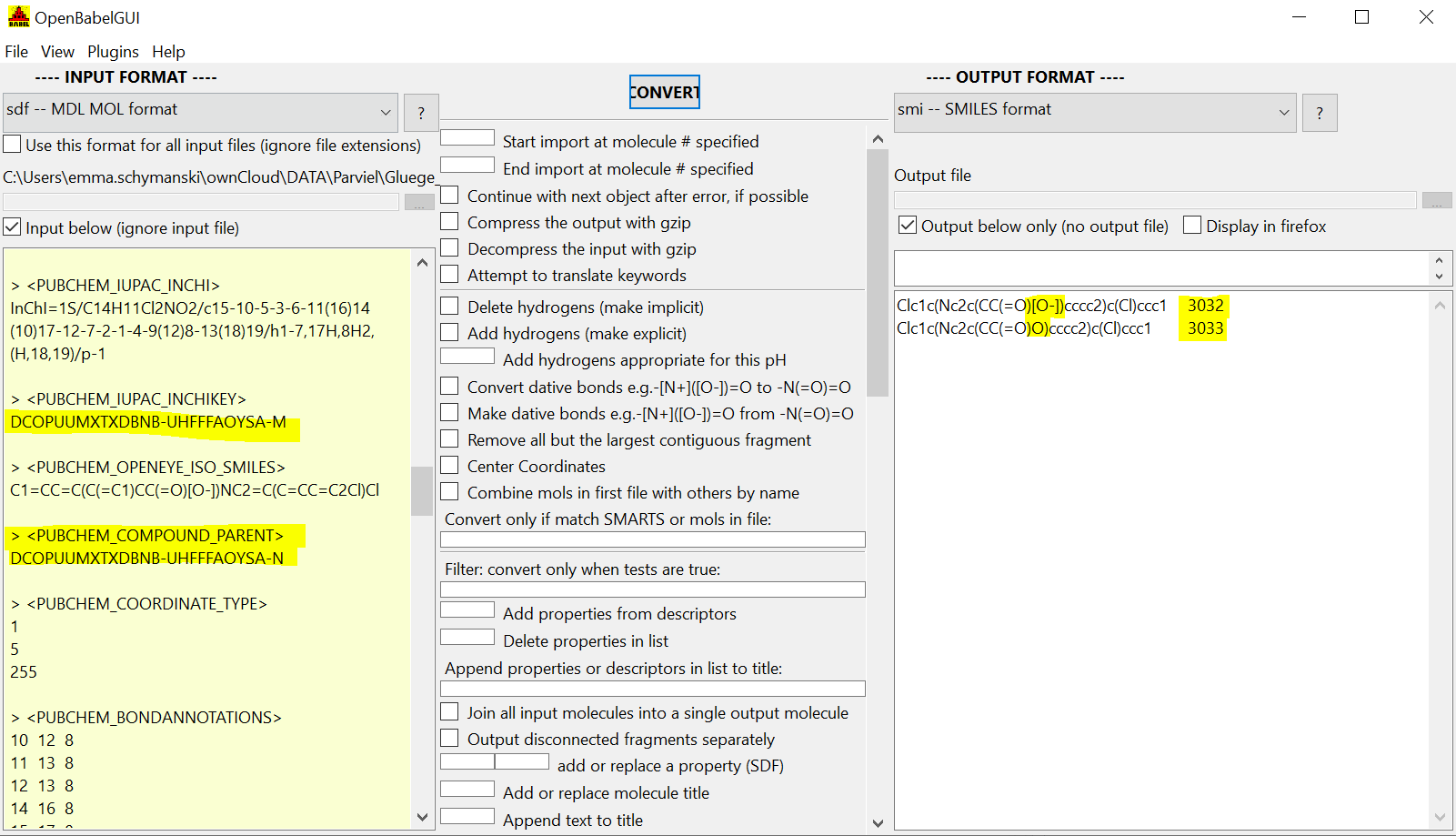
Simpler case DCOPUUMXTXDBNB-UHFFFAOYSA-M.sdf:
- charged entry in
- charged and neutral forms back, with parent InChIKey pointing to neutral form in both entries
Another case (known tautomer issue): QTXVAVXCBMYBJW-UHFFFAOYSA-N.sdf
- only one entry back, the InChIKey doesn't match because it's PubChem's preferred tautomer of warfarin
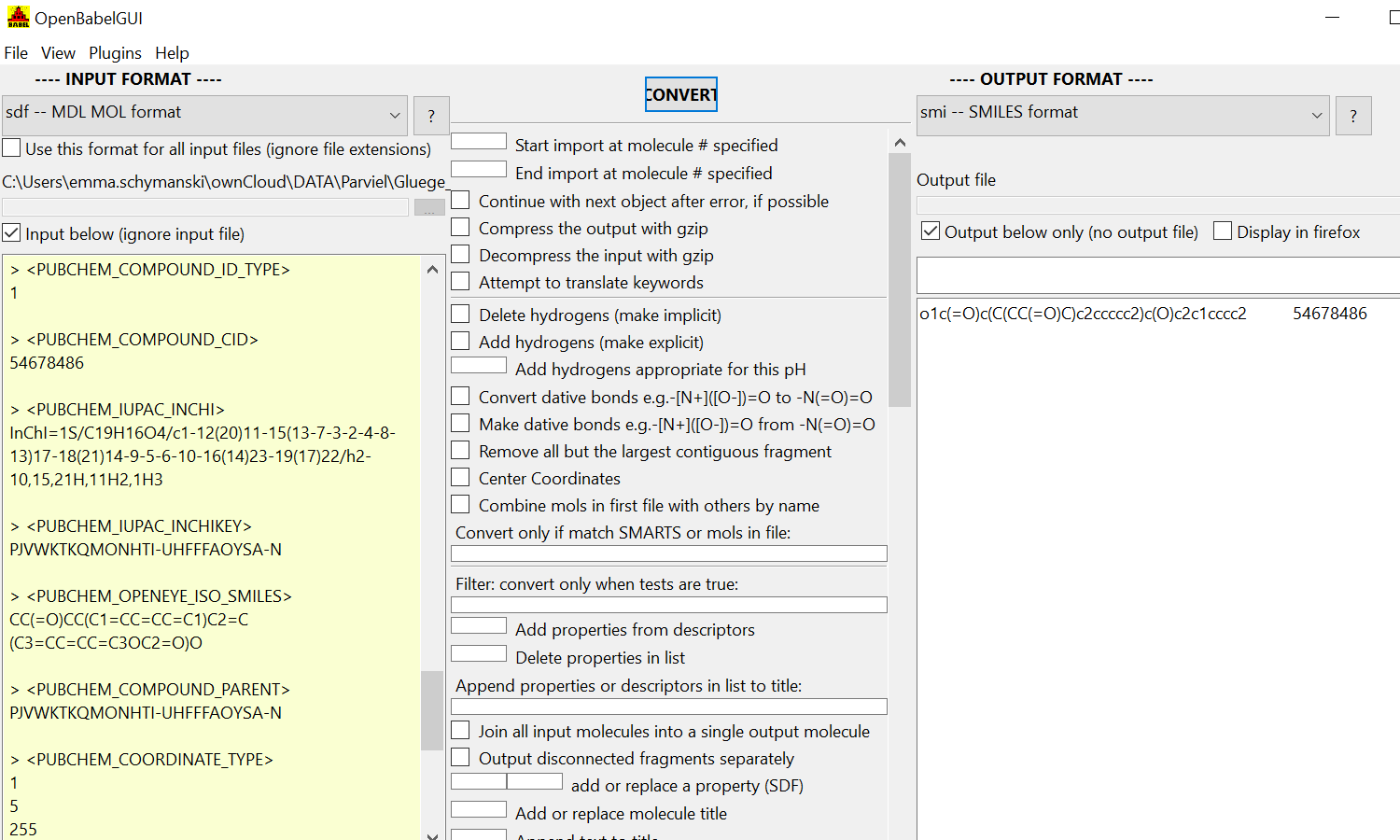
(see https://pubchem.ncbi.nlm.nih.gov/#query=QTXVAVXCBMYBJW-UHFFFAOYSA-N vs https://pubchem.ncbi.nlm.nih.gov/#query=PJVWKTKQMONHTI-UHFFFAOYSA-N
There are several different cases in the test set ... happy to explain more on Monday!
PS: seems I can't post issues in https://github.com/ZeroPM-H2020/pcapi hence posting here...